GRACE normals processing¶
[1]:
# Imports
from gfeatpy.gravity import SphericalHarmonicsCovariance, EquivalentWaterHeight
from gfeatpy import plotting
import matplotlib.pyplot as plt
data_root = "../../../data"
Non-resonant case¶
[2]:
# Load monthly normal matrix
monthly_error = SphericalHarmonicsCovariance(96)
monthly_error.from_normal(f"{data_root}/gravity/monthly_normals/ITSG-Grace2018_n96_2007-03.snx", scaling_factor = 0.9330416388987962) # EXPENSIVE!! ~1 min
[2]:
<gfeatpy.gravity.SphericalHarmonicsCovariance at 0x7502ded06170>
[3]:
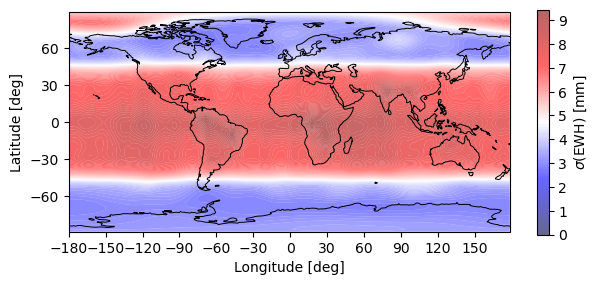
# Plot EWH commission error
plotting.synthesis(monthly_error, 180, 90, EquivalentWaterHeight(0))
plt.show()
Warning: Number of longitude nodes increased to 192 reach full SH spectrum during FFT synthesis over parallels.
[4]:
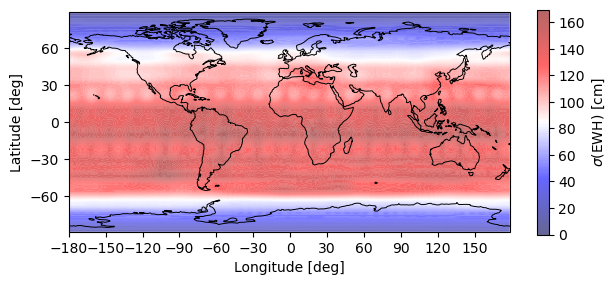
# Plot EWH + Gaussian spatial smoothing filter r=750 km
plotting.synthesis(monthly_error, 180, 90, EquivalentWaterHeight(750e3))
plt.show()
Warning: Number of longitude nodes increased to 192 reach full SH spectrum during FFT synthesis over parallels.
Resonant case¶
[5]:
# Load monthly normal matrix
monthly_error = SphericalHarmonicsCovariance(96)
monthly_error.from_normal(f"{data_root}/gravity/monthly_normals/ITSG-Grace2018_n96_2004-09.snx", scaling_factor = 1.0) # EXPENSIVE!! ~1 min # scale factor incorrect!!
[5]:
<gfeatpy.gravity.SphericalHarmonicsCovariance at 0x7502ddc26cb0>
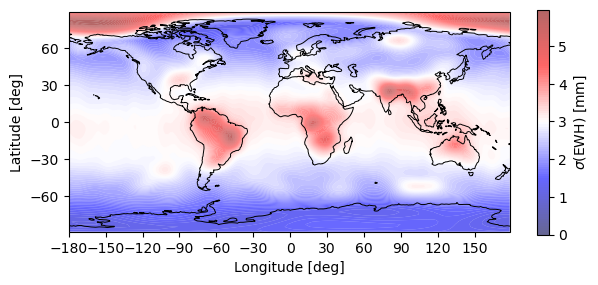
[6]:
# Plot EWH
plotting.synthesis(monthly_error, 180, 90, EquivalentWaterHeight(0))
plt.show()
Warning: Number of longitude nodes increased to 192 reach full SH spectrum during FFT synthesis over parallels.
[7]:
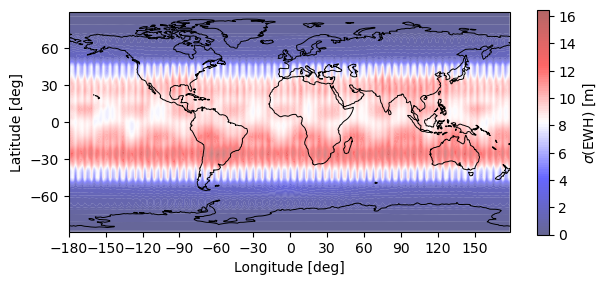
# Plot EWH + Gaussian spatial smoothing filter r=750 km
plotting.synthesis(monthly_error, 180, 90, EquivalentWaterHeight(750e3))
plt.show()
Warning: Number of longitude nodes increased to 192 reach full SH spectrum during FFT synthesis over parallels.